Carbon atoms and molecules containing carbon-carbon bonds are central to all living organisms on Earth. For example, they are the backbone of DNA, RNA, proteins, plants, animals, humans, and much more. Even so, is it possible for life to be centered around an element other than carbon? The periodic table, introduced in the 1860s, orders elements by size and chemical properties. Elements in the same column are predicted to exhibit similar Chemical properties. Silicon should be the most chemically similar to carbon since it is located directly below carbon in the periodic table. They both have four valence electrons in their outer shell so both are capable of bonding four groups to form tetrahedral structures.
However, there are a few key differences between the
bonds that carbón and silicon can make. Carbon-carbon bonds are slightly
stronger than silicon-silicon bonds due to the larger size of silicon atoms.
Carbon is limited by the octet rule to eight valence electrons and is generally
limited to four bonds. Silicon is not constrained by the octet rule and can
have more than eight valence electrons; it prefers four bonds but easily makes
stable molecules where it has five or even six bonds to other atoms.
The ability to form more bonds should allow silicon to
produce a wider array of complex molecular structures than carbon.
Additionally, silicon predominates over carbon in rocky planets like Earth.
Silicon makes up 14% of Earth by mass whereas carbon makes up less than 0.1%.
Given the chemical promiscuity and abundance of silicon can we envision life
forms based on silicon instead of carbon?
The question of silicon based life isn’t new. Many
have speculated on silicon-based life forms and Isaac Asimov’s biochemical
training and expansive vision led him to assess the chemical features of carbon
relative to silicon. Both elements form highly stable oxides with molecular
oxygen: carbon forms carbon dioxide (CO2) and silicon forms silica (SinO2n).
The physical properties of these two compounds differ dramatically. Carbon
dioxide is readily processed; it is a gas that readily dissolves in and reacts
with water. In contrast, silica readily forms refractory silicate salts. Silica
and silicates are the main component of rock; they are insoluble in water and
unreactive. Furthermore, the higher stability of silica relative to carbon
dioxide makes it harder to reduce to form useful Si-H or Si-Si bonds. Life on
our planet is ultimately driven by biochemical reduction of CO2 to make
molecules composed of C-H and C-C bonds.
Life is also dependent on a host of other biochemical
processes such as replication, adaptation, and metabolism. These processes are
dependent on interactions involving flat rings that are one atom thick. The
flat shape of such molecular fragments is dependent upon the ability to form
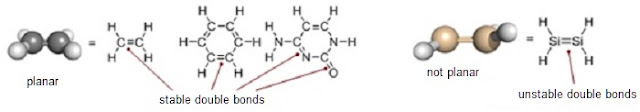
intrinsically stable, flat, double bonds (Figure 1). Carbon atoms can do it;
nitrogen atoms can do it; oxygen atoms can do it; and they can combine to form
stable carbon-nitrogen, carbon-oxygen, and nitrogen-oxygen doublé bonds.
Silicon atoms cannot form stable double bonds with each other or with other atoms.
Silicon-silicon double bonds are weaker than a typical carbon-carbon doublé bond
(14-24 kcal/mol vs >48 kcal/mol) and require extremely large groups on
silicon to prevent spontaneous reactions with oxygen or water. Silicon-silicon
double bonds are not planar because the silicon atoms prefer pyramidal
geometries.
Figure 1: Examples of stable double bonds involving carbon and an example of an unstable silicon-silicon double bond.
The ability of carbon to form stable flat molecules
with other carbon atoms makes it a powerful atom. In one of its purest forms –
graphite – carbon atoms are bonded with each other to form flat sheets. Each
carbon atom in graphite is attached to three other carbon atoms, forming a
two-dimensional hexagonal array that resembles a honeycomb structure (Figure 2,
left). In a simplified representation, each carbon atom in graphite is doubly
bonded to one other carbon atom. These double bonds are also known as pi bonds.
The two-dimensional sheets of carbon atoms in graphite stack on top of each
other. When the sheets stack, the pi bonds from one sheet interact with pi bonds
on sheets above and below.
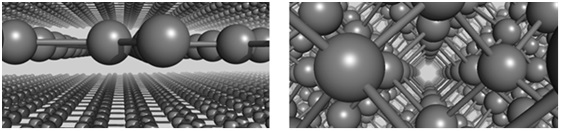
Figure 2: Left: In graphite, carbon atoms are arranged in stacked sheets. Right: In diamond, carbon atoms are arranged in a 3-dimensional lattice.
Those interactions are referred to as pi stacking interactions,
which are stabilizing; they make the aggregate of sheets stronger than the
individual sheets. Diamond is another pure form of carbon in which each carbon
atom is bonded to four other carbon atoms forming a rigid three-dimensional
lattice (Figure 2, right). Under normal conditions, graphite is slightly more
stable than diamond because of the favorable pi stacking interactions between
the sheets. The energy of these sheet like interactions in graphite is
approximately 1.4 kcal/mol, making stacked sheets of carbón ten times more
stable than isolated sheets.
Pi stacking is equally possible when the flat
molecules contain double bonds between carbon, nitrogen, and oxygen in various
combinations. The molecules of life are completely dependent on these pi stacking
interactions between atomically flat components. For example, in DNA, pi
stacking of subunits called “bases” is essential for self-assembly into the
iconic double helix (Figure 3, left). The DNA bases are flat rings composed of
carbon, nitrogen, and oxygen atoms. The pi stacking interactions between DNA
bases are essential for the long term stability of DNA and its function as a
repository for genetic information. The biological molecule RNA also contains
bases and is widely recognized as the molecular precursor to DNA in the genesis
of life. Not surprisingly, similar pi stacking interactions are evident in the
chemical structure of RNA (Figure 3, center). Finally, inspection of proteins
reveals the importance of pi stacking interactions both within the folded
structure of proteins and between proteins and their reacting partners (Figure
3, right).
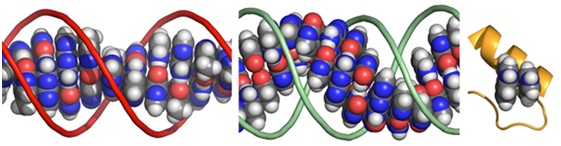
Figure 3: Pi-stacking interactions between flat bases in the molecules of life. Left: flat bases stack up in DNA; Middle: flat bases stack up in RNA; Right: protein side chains stack in a metabolic protein enzyme.
Carbon can form stable flat double bonds and
atomically flat rings that are necessary for the molecular recognition
processes responsible for life. Silicon does not form stable double bonds with
any atom: nitrogen, carbon, oxygen or even itself. Since silicon cannot form
stable double bonds, it cannot form atomically flat structures capable of
stacking. A biochemical universe based on silicon would lack the capacity for
face-to-face molecular recognition through pi-stacking, much like a warehouse containing
irregular containers that could not be stacked.
A silicon-based biochemical system faces a serious
design challenge that is not easily overcome. Even if all environmental
conditions were met to allow for silicon containing bonds to the main
constituent of biochemical processes, silicon will always fall short. In a
world where life depends on molecular processes involving atomically flat
structures, silicon will never be able to stack up.
Bibliography:
1. a. Asimov, I. “Big Brother” The Magazine of Fantasy and Science Fiction.
September, 1982. b. Asimov, I. “V. Big Brother” X Stands for Unknown. Doubleday: New York, 1984. p. 61-71.
2. a. Asimov, I. “Bread and Stone” The Magazine of Fantasy and Science Fiction.
October, 1982. b. Asimov, I. “VI. Bread and Stone” X Stands for Unknown. Doubleday: New York, 1984. p. 72-82.
3. a. Asimov, I. “VII. A Difference of an E” The Magazine of Fantasy and Science Fiction.
November, 1982. b. Asimov, I. “VII. Big Brother” X Stands for Unknown. Doubleday: New York, 1984. p. 83-94.
4. a. Asimov, I. “Silicon Life After All” The Magazine of Fantasy and Science Fiction.
December, 1982. b. Asimov, I. “VIII. Silicon Life After All” X Stands for Unknown. Doubleday: New
York, 1984. p. 95-108.
5. Iwamoto, T.; Ishida, S.
“Multiple Bonds with Silicon: Recent Advances in Synthesis, Structure, and
Functions of Stable Disilenes” in Functional
Molecular Silicon Compounds II. Scheschkewitz, D., Ed. Structure and
Bonding, Vol 156. Springer: Switzerland, 2013. pp. 125-202.
6. Zacharia, Renju “Chapter 4. Energetics of
interlayer binding in graphite.” in Desorption
of Gases from Graphitic and Porous Carbon Surfaces. Dissertation. Freie
Universtitat Berlin. 2004.
7. Girifalco, L. A.; Lad, R. A. Lad, J. Chem. Phys., 1956, 25, 693.
8. Winter, N. W.; Ree, F. H.
“Stability of the Graphite and Diamond Phases of Finite Carbon Clusters” Detonation Symposium Snowmass, CO August
30 – September 4,1998.
David L. Van Vranken Vanessa Arredondo
Professor of Chemistry Ph.D. Candidate in Chemistry**
University
of California, Irvine
** (Ph.D. 2019)
Vanessa Arredondo
** (Ph.D. 2019)

No hay comentarios:
Publicar un comentario